EEG dataset
A public data set provided by Mumtaz et al. was utilized to evaluate the proposed method30. A total of 64 subjects participated in the experiment, including 34 MDD subjects (17 men, average age 40.3 ± 12.9) and 30 normal control (NC) subjects (21 male, average age 38.23 ± 15.64), all participants are from Hospital Universiti Sains Malaysia (HUSM). MDD participants with psychotic symptoms, alcoholism, smoking and epilepsy were excluded from the study. The healthy control group was also excluded for possible mental or physical illness. All participants signed an informed consent form and were informed of the details of the trial. The experiment was designed in accordance with the Helsinki Declaration and approved by the HUSM Ethics Committee. The 19-channel EEG cap was used for EEG recording, and the electrodes were placed in the international standard 10–20 system, including Fp1, F3, C3, P3, O1, F7, T3, T5, Fz, Fp2, F4, C4, P4, O2, F8, T4, T6, Cz and Pz, as shown in Fig. 2, where A1 and A2 were the reference electrodes.
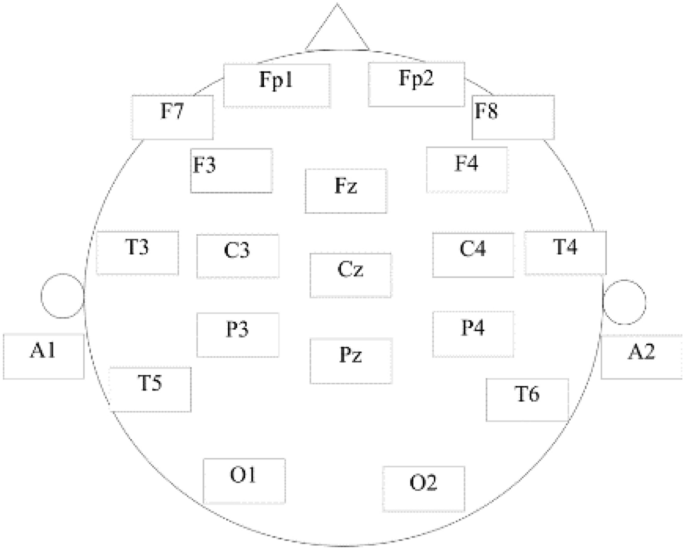
Electrode placement position.
The resting-state EEG signals were acquired from MDD and HC subjects in the eye-opened (EO) condition. These signals are sampled at 256 HZ and filtered with a 0.5 Hz to 50 Hz bandpass filter and an additional 50 Hz notch filter. The artifacts were then removed using EEGLAB. Subjects 14 and 25 in the NC group and 7, 8, 12, and 34 in the MDD group were excluded because the EEG was less than 4 min after processing. Therefore, the data used in this study came from 58 subjects, including 28 NC and 30 MDD. Meanwhile, the EEG of the middle four minutes was intercepted to reduce the effect of noise at both ends. In order to improve the sample size, each subject’s data were segmented according to 10 s, and finally, a total of 1392 (58 × 24) samples were obtained.
EEG features
The sampling frequency of EEG signals is 256HZ, so six layers of wavelet transform are designed to extract delta, theta, alpha, beta, and gamma bands. On this basis, the Hjorth parameter, approximate entropy (ApEn), power spectral density (PSD), fractal dimension (FD), and CO complexity were extracted from each frequency band. Finally, the features of all channels and frequency bands are combined as feature vectors. Table 1 shows the specific information of EEG features. The number of signal channels is 19, and the signal of each channel is decomposed into five frequency bands, so the dimension of the feature is 95 (19 × 5). The Hjorth parameter contains three perspectives, so its dimension is 285 (95 × 3).
Ablation study
To verify the effectiveness of the proposed method, an ablation experiment was conducted, and the experimental setup is shown in Table 2. MQSVM1, MQSVM2, and MPSVM are hybrid feature selection methods, FS-QPSO is Wrapper feature selection, and FS-MIC is Filter feature selection. Five-fold cross-validation was used in the experiments. Five-fold cross-validation is to divide the data into five parts, each time extracting one part (not repeated) as the verification set, the remaining four parts as the training set. The training set is used to train the model. Also, three-fold cross-validation is used in the model training, that is, the k value of the proposed method is set to 3. The trained model is then applied to the validation set and the experimental results are obtained. The above operation is repeated five times and the results are averaged to obtain the final result. MPSVM uses PSO to optimize feature subsets and classifier parameters. The two acceleration coefficients of the PSO are set to 2, the inertia weight to 1, and the number of iterations and particles to 200. Other methods use QPSO to optimize feature subsets or classifier parameters. The number of iterations and particles in the QPSO is set to 200. FS-MIC uses a forward search strategy to find the optimal subset. All experiments were performed on Intel(R) Xeon(R) CPU, 2.3 GHz, 128 GB RAM.
Figure 3 shows the optimization process of MQSVM1 and MQSVM2 in the second stage. It can be seen that the fitness values of MQSVM1 and MQSVM increase with the number of iterations. Meanwhile, the fitness value reaches the convergence value before the end of the iteration, indicating that the particle number and iteration number settings are valid. The convergence value of MQSVM2 is lower than that of MQSVM1 for different EEG features because PSO tends to get stuck in local optima, while QPSO can jump out of local optima. The above analysis shows that the performance of the proposed method is higher when QPSO is used in the second stage.
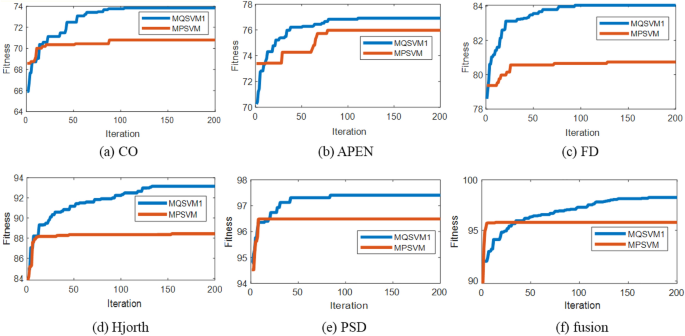
The change curve of fitness value in the second stage.
Table 3 shows the running times of all methods. It can be found that the running time of all methods increases with the feature dimension. The running time of FS-MIC increases significantly with the feature dimension, since FS-MIC employs a forward search strategy, which requires the QPSO to optimize the classifier parameters each time a feature is accepted. The running time of MQSVM1 gently increases with the feature dimension, since MQSVM1 uses QPSO to optimize the classifier parameters while optimizing a subset of features. When the feature dimension is low (CO, APEN, FD), the running time of FS-MIC is small, which is due to the low computational complexity of the Filter method. When the feature dimension is high (fusion), the running time of MQSVM1 is small, indicating that the proposed method is suitable for high-dimensional data sets. In terms of average running time, FS-QPSO has the longest running time, while the other four methods have almost no difference in running time, indicating that the Wrapper method has the highest computational complexity. In particular, the running time of FS-QPSO is very long in high-dimensional features, which is not favorable for practical applications. The above analysis shows that the proposed method is not computationally complex and can be used for EEG feature selection.
Table 4 shows the dimensions of the feature subsets. It can be seen that the feature subset dimension after feature selection is significantly lower than the original feature dimension. The dimension of feature subsets obtained by three hybrid feature selection methods (MQSVM1, MQSVM2, MPSVM) is significantly lower than that of the Wrapper and the Filter feature selection methods, which indicates that the hybrid feature selection method has advantages in feature selection. Among all EEG features, the dimensionality of the feature subset obtained by the proposed method is significantly lower than that of the other two hybrid feature selection methods, indicating that the proposed improved strategy is effective. The dimensionality of the feature subset of MQSVM1 is significantly lower than that of MQSVM2, which indicates that the fitness function constructed in this paper using the dimensionality of the feature subset and the classification accuracy is effective. The dimension of the feature subset of MQSVM1 is significantly lower than that of MPSVM, indicating that PSO is more likely to get stuck in the local optimal solution when optimizing the feature subset. At the same time, QPSO can jump out of the local optimal solution, so the dimension of the feature subset obtained by MQSVM1 is low. The above analysis shows that the dimension of the feature subset obtained by the proposed method is lower than that of the other methods, which is beneficial for reducing the computational complexity of the proposed method.
Table 5 shows the classification accuracy of EEG. It can be found that the classification accuracy of FS-QPSO is significantly higher than the other methods because the Wrapper method can evaluate the overall performance of feature subsets, which is beneficial for improving the classification accuracy of feature subsets. The classification accuracy of MQSVM1 is significantly higher than that of FS-MIC, which indicates that the proposed method can combine the advantages of the Wrapper method and improve the classification accuracy of the proposed method. The classification accuracy of MQSVM1 is higher than that of MPSVM, which indicates that QPSO has more optimization power than PSO, and therefore it is more advantageous to use QPSO for feature optimization.
The weighted values of feature subset dimension and classification accuracy were calculated according to Eq. (28), and the results are shown in Table 6. The weighted values synthesize the dimensionality and classification accuracy of a subset of features, which facilitates direct comparison and analysis of the performance of different methods. The larger the weighted value, the better the performance of the feature subset obtained by the method. It can be found that the weighted value of FS-MIC is significantly lower than that of the other methods, indicating that the feature subset obtained by the Filter feature selection method performs relatively poorly. The weighted values of MQSVM1, MQSVM2, and FS-QPSO are higher, which indicates that the feature subsets obtained by the hybrid feature selection method and the Wrapper feature selection method have better performance. The weighted value of MPSVM is significantly lower than that of the other two hybrid feature selection methods, indicating that QPSO performs significantly better than PSO in feature subset optimization. In several EEG features (CO, APEN, Hjorth, PSD, fusion), MQSVM1 was weighted higher than other methods. Considering the average values, the weighted value of MQSVM1 is the largest, indicating that the feature subset obtained by the proposed method performs the best.
The proposed method runs faster when only MIC is used for feature selection. However, since the Filter feature selection method uses a forward search approach, the running time in high-dimensional datasets is extended when the classifier needs to optimize parameters. Therefore, the running time of Filter feature selection methods combined with some classifiers that do not require parameter optimization will be significantly reduced. In addition, the feature dimension and accuracy obtained by the Filter feature selection method are lower than those of Wrapper and hybrid feature selection methods because the feature subset obtained by the Filter feature selection method lacks the overall evaluation of the feature subset, so the performance of the feature subset obtained by the Filter feature selection method is not high. FS-QPSO achieves the highest classification accuracy, but its running time is longer than Filter and hybrid feature selection methods due to the ample search space. When the feature dimension is higher (fusion), the running time increases significantly, reaching 1214 min, which makes the method difficult in practical application. Therefore, although the feature subset obtained by the Wrapper feature selection method performs well, the computational complexity is too high, and it is not easy to apply it directly to high-dimensional features. MQSVM1 combines the advantages of Filter and Wrapper feature selection methods, and its runtime is significantly lower than FS-QPSO. The proposed method optimizes the parameters of the classifier while optimizing a subset of features in the second stage, which reduces the computational complexity of the method in high-dimensional features to some extent. Therefore, the running time of the proposed method is significantly slower than that of FS-QPSO as the dimension of EEG features increases. There are differences between MQSVM1 and MQSVM2 in terms of fitness function. MQSVM1 uses a fitness function constructed from feature subset dimension and classification accuracy, and MQSVM2 uses a fitness function constructed from classification accuracy. Experimental results show that the feature subset dimension of MQSVM1 is significantly lower than that of MQSVM2, and the classification accuracy of MQSVM1 is also high, which indicates that the fitness function designed in this paper can effectively reduce the dimension of the low feature subset and achieve high classification accuracy. MQSVM1 and MPSVM have different optimization algorithms. MQSVM1 uses QPSO to optimize feature subsets, and MPSVM uses PSO to optimize feature subsets. Experimental results show that the classification accuracy of MQSVM1 is significantly higher than that of MPSVM, which indicates that QPSO has a more robust search capability and can significantly reduce the probability of the method falling into local optimality. Therefore, it is efficient to use QPSO in the second stage to optimize the feature subset. In terms of weighted values, MQSVM1 achieves significantly higher values than the other methods, which verifies the effectiveness of the proposed improved strategy.
Comparison with existing methods
To thoroughly verify the superiority of the proposed method, we compared the proposed method with other classical methods, including EEG feature selection methods and other domain feature selection methods. These include the MRMR EEG feature selection method adopted by Cai et al. in 201811, the Fscore-based EEG feature selection method adopted by Wu et al. in 201824, and the RFINCA EEG feature selection method adopted by Tuncer et al. in 202112, the ReliefF-QPSO feature selection method adopted by Xue Rui et al. in 202015, and the Fscore-MIC feature selection method adopted by Zhao Ling et al. in 202113.
The experimental setup is the same as in Section “Ablation study”. The experiments are performed with five-fold cross-validation, the model is trained with three-fold cross-validation, and the number of particles and iterations of the QPSO is set to 200. The number of neighbors for ReliefF-QPSO and RFINCA is set to six. For the other methods, no other parameters need to be set. MRMR, Fscore, FSCOre-MIC, and RFINCA use a forward search strategy to find the optimal subset.
Table 7 shows the running times of the different methods. It can be found that as the feature dimension increases, the running time of all methods increases. MQSVM and Relief-QPSO are hybrid feature selection methods, but the running time of MQSVM is significantly lower than that of Relief-QPSO, mainly because ReliefF only deals with irrelevant features when performing feature selection. The proposed method uses MIC to deal with irrelevant and redundant features, which reduces the search space in the second stage and hence the computational complexity of the method. The remaining four methods are Filter feature selection methods, which have significantly lower running times than hybrid feature selection methods in low-dimensional features. However, the running time of the Filter feature selection method increases significantly in high-dimensional features, mainly because the Filter method uses a forward search method to find the optimal subset of features. Compared with existing methods, the running time of the proposed method grows more slowly with the increase of feature dimension, mainly because in the second stage, the proposed method optimizes the parameters of the classifier while optimizing the feature subset. The average runtime of MQSVM is 227 min, which is not a significant increase over existing methods. The above analysis shows that the computational complexity of the proposed method is not very high compared to the existing methods, and it increases slowly in high-dimensional features.
Table 8 shows the dimensions of the feature subsets. It can be found that the dimensionality of the feature subsets obtained by all methods is significantly reduced. The dimension of the feature subset obtained by Relief-QPSO is significantly higher than that obtained by the other methods. In high-dimensional features (Hjorth, fusion), the performance of Relief-QPSO is significantly reduced, which is because the fitness function of Relief-QPSO does not consider the influence of the feature subset dimension, and the ReliefF only processes irrelevant features, resulting in a large search space in the second stage. The dimension of the feature subset obtained by MQSVM is significantly lower than that of Relief-QPSO because MQSVM uses MIC to process irrelevant and redundant features, which reduces the search space in the second stage. In terms of the mean value of the results, the proposed method has the smallest value, indicating that the dimension of the feature subset obtained by the proposed method is lower than that of the existing methods.
Table 9 shows the classification accuracy. It can be found that the classification accuracy of MQSVM is not much different from the existing methods, so it is difficult to judge the performance of these methods from the classification accuracy alone.
The weighted sum of the dimension and classification accuracy of the feature subset was calculated according to Eq. (28) to analyze the performance of different methods. The results are shown in Table 10. It can be found that MQSVM has significantly higher weighting values on multiple EEG features (CO, FD, Hjorth, PSD, fusion) than existing methods. At the same time, MQSVM has the largest value in terms of the mean value of the results. The above analysis shows that the feature subset obtained by the proposed method has higher classification accuracy and lower feature dimensionality than the existing methods.
The average running time of the proposed method is 227 min, which is significantly lower than the existing hybrid feature selection method, indicating that the computational complexity of the proposed method is not high. At the same time, the running time of the proposed method gently increases with the feature dimension, which indicates that the proposed method performs well on high-dimensional datasets. The dimension of the feature subset obtained by the proposed method is lower than that of the existing method, and the classification accuracy of the feature subset obtained by the proposed method is not much different from that of the existing method, which indicates that the feature subset obtained by the proposed method not only has a lower dimension but also has high classification accuracy. The weighted sum of the dimensionality and classification accuracy is computed, which facilitates a direct comparison of the performance of the feature subsets obtained by different methods. The weighted value of the proposed method is higher than that of the existing methods, which indicates that the performance of the feature subset obtained by the proposed method is better than that of the existing methods. At the same time, the weighted values of the proposed method are larger than those of the existing methods in many EEG features, which indicates that the performance of the proposed method is relatively stable across different EEG features. The above analysis shows that the feature subsets obtained by the proposed method have low dimensionality, high classification accuracy, and low computational complexity compared to the existing methods, which validates the effectiveness of the proposed method.
Analyze the robustness of the proposed method
To further validate the robustness of the proposed method, the UCI dataset is used for experimental validation. The details of the datasets are given in Table 11. The UCI dataset is divided into a training set and a validation set; the training set is 75 percent, and the validation set is 25 percent. The parameter settings for the algorithm are the same as those used for the experiments in Section “Comparison with existing methods”.
The weighted sum of the dimension and classification accuracy of the feature subset is calculated according to Eq. (28), and the results are shown in Table 12. It can be found that the weighted value of the proposed method is significantly higher than that of the existing method on multiple datasets (segment, dermatology, Parkinson, CNAE, QSAR) and has obvious superiority on high-dimensional datasets (Parkinson, CNAE, QSAR). By averaging the results over all datasets, it can be found that the mean value of MQSVM is significantly higher than that of the existing methods. The larger the weighted value, the better the overall performance of the feature subset. The above analysis shows that the proposed method has stable performance on datasets with different dimensions, and the extracted feature subsets perform well, which indicates that the proposed method is robust and can be used for feature extraction on other data.